
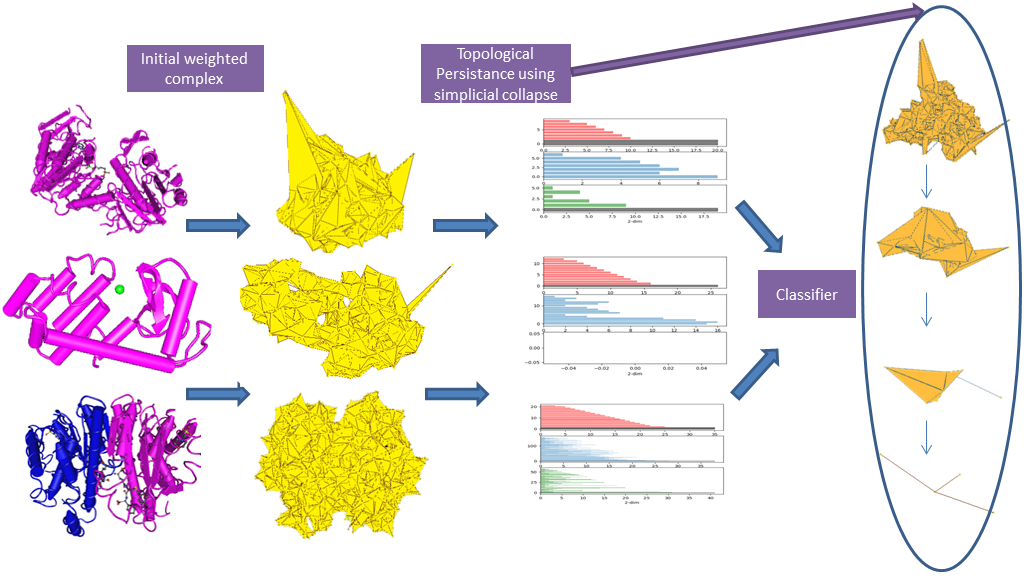
Automated annotation and analysis of protein molecules have long been a topic of interest due to immediate applications in medicine and drug design. In this work, we propose a topology based, fast, scalable, and parameter-free technique to generate protein signatures. We build an initial simplicial complex using information about the protein’s constituent atoms, including radius and existing chemical bonds, to model the hierarchical structure of the molecule. Simplicial collapse is used to construct a filtration which we use to compute persistent homology. This information constitutes our signature for the protein. In addition, we demonstrate that this technique scales well to large proteins. Our method shows sizable time and memory improvements compared to other topology based approaches. We use the signature to train a protein domain classifier. Finally, we compare this classifier against models built from state-of-the-art structure- based protein signatures on standard datasets to achieve a substantial improvement in accuracy. This work is supported by NSF grants CCF-1318595, CCF-1526513, and CCF-1733798
OTHER PUBLICATIONS OF OUR RESEARCH GROUP
Disclaimer: We do not intend to be responsible for the maintenance of the software.
Copyright: Jyamiti group at the Ohio State University. No commercial use of the software is permitted without proper license.